τ-Random Acceleration Molecular Dynamics (τRAMD)
Introduction
An extension to the RAMD method for calculating relative residence times (τ) for a set of small molecules bound to a target protein.
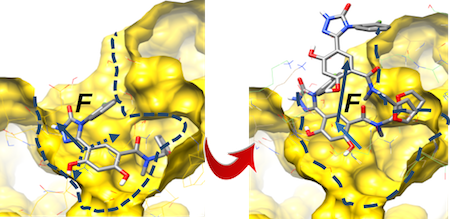 τRAMD extends the RAMD method to accelerate the egress of a set of small molecules from a target receptor. Slower dissociating molecules take longer to leave the binding pocket, or require application of a stronger force to exit within a specified simulation time. Similarly, molecules with higher dissociation rates make their exit the binding pocket more quickly, and require comparatively smaller forces for egress to occur within a defined simulation time. Thus, τRAMD may provide a computationally efficient approach to obtain relative estimates of dissociation rate constants.
τRAMD extends the RAMD method to accelerate the egress of a set of small molecules from a target receptor. Slower dissociating molecules take longer to leave the binding pocket, or require application of a stronger force to exit within a specified simulation time. Similarly, molecules with higher dissociation rates make their exit the binding pocket more quickly, and require comparatively smaller forces for egress to occur within a defined simulation time. Thus, τRAMD may provide a computationally efficient approach to obtain relative estimates of dissociation rate constants.
Software and instructions for running RAMD and τRAMD simulations in NAMD can be found here.
and here (Tutorial).
Software and instructions for running RAMD and τRAMD simulations in Gromacs can be found here (Gromacs-RAMD).
and here (Tutorial).
References:
1. Kokh DB et. al. Estimation of Drug-Target Residence Times by τ-Random Acceleration Molecular Dynamics Simulations. J. Chem. Theory Comput. 2018; 14(7): 3859-3869 DOI: 10.1021/acs.jctc.8b00230
2. Kokh DB et. al. A workflow for exploring ligand dissociation from a macromolecule: Efficient random acceleration molecular dynamics simulation and interaction fingerprint analysis of ligand trajectories, J. Chem. Phys. 153, 125102 (2020); DOI: 10.1063/5.0019088
3. Maximova E. et. al. Protein–Ligand Dissociation Rate Constant from All-Atom Simulation, The Journal of Physical Chemistry Letters, Vol 12 Issue 43 (2021);
4. Yiwei Ding, et. al. A theoretical framework for random acceleration molecular dynamics simulations. The Journal of Chemical Physics 2026, 164 (6) https://doi.org/10.1063/5.0309358
Example Cases
For examples of previously performed studies in which τ-Random Acceleration Molecular Dynamics (τRAMD) was the primary method used, see the following example cases:
- Investigating the unbinding mechanisms and kinetics of MmpL3 inhibitors: A computational study.
- Role of UDP-N-acetylmuramic acid in the regulation of MurA activity revealed by molecular dynamics simulations.
- Characterization of the Bottlenecks and Pathways for Inhibitor Dissociation from [NiFe] Hydrogenase.
- Structure-kinetic relationship reveals the mechanism of selectivity of FAK inhibitors over PYK2.
- G Protein-Coupled Receptor-Ligand Dissociation Rates and Mechanisms from τRAMD Simulations.
- Ligand unbinding mechanisms and kinetics for T4 lysozyme mutants from τRAMD simulations.
- Protein-Ligand Dissociation Rate Constant from All-Atom Simulation.
- Machine Learning Analysis of τRAMD Trajectories to Decipher Molecular Determinants of Drug-Target Residence Times
- Estimation of drug-target residence times by τ -random acceleration molecular dynamics simulations.
Tutorials
The folowing tutorials describe the use of τ-Random Acceleration Molecular Dynamics (τRAMD):