Metadynamics
Introduction
Metadynamics is an enhanced sampling method and is informally described as "filling the free energy wells with computational sand".
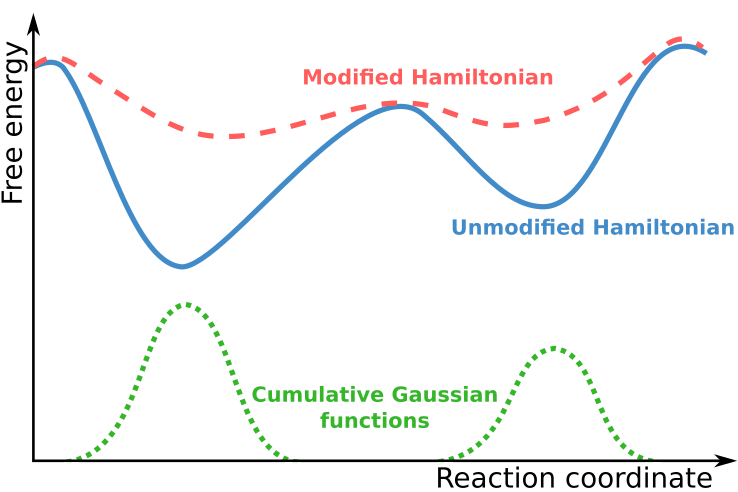
In Metadynamics simulations, the system is described using a set of collective variables that define transitions along a reaction coordinate. During the simulation, the location of the system in this collective variable space is determined, and positive biasing Gaussian functions are added to the overall potential, thus modifying the Hamiltonian of the system.
\(H = T + V + \sum V_\textrm{GAUSS}\)
The accumulation of these Gaussian functions in well sampled regions of the collective variable space results in a flattening of the free energy surface, thereby allowing the system to more easily traverse regions of collective variable space that correspond to free energy maxima when simulating the unmodified Hamiltonian. This allows the simulation to explore the full energy landscape. With knowledge of the sampling of the modified Hamiltonian, and the deposited Gaussian functions, it is possible to recover the free energy surface of the unmodified Hamiltonian.
Example Cases
For examples of previously performed studies in which Metadynamics was the primary method used, see the following example cases:
- Computational Estimation of Residence Time on Roniciclib and Its Derivatives against CDK2: Extending the Use of Classical and Enhanced Molecular Dynamics Simulations.
- Prediction of Threonine-Tyrosine Kinase Receptor-Ligand Unbinding Kinetics with Multiscale Milestoning and Metadynamics.
- Unbinding Kinetics of Muscarinic M3 Receptor Antagonists Explained by Metadynamics Simulations.
- Accuracy of Molecular Simulation-Based Predictions of koff Values: A Metadynamics Study.
- A combination of machine learning and infrequent metadynamics to efficiently predict kinetic rates, transition states, and molecular determinants of drug dissociation from G protein-coupled receptors.
- An Integrated Markov State Model and Path Metadynamics Approach to Characterize Drug Binding Processes.
- Unbinding Dynamics of Non-Nucleoside Inhibitors from HIV-1 Reverse Transcriptase.
- Exhaustive Search of Ligand Binding Pathways via Volume-Based Metadynamics.
- Accelerating the Calculation of Protein-Ligand Binding Free Energy and Residence Times Using Dynamically Optimized Collective Variables.
- Ligand Binding, Unbinding, and Allosteric Effects: Deciphering Small-Molecule Modulation of HSP90.
- Chasing the Full Free Energy Landscape of Neuroreceptor/Ligand Unbinding by Metadynamics Simulations.
- Can One Trust Kinetic and Thermodynamic Observables from Biased Metadynamics Simulations?: Detailed Quantitative Benchmarks on Millimolar Drug Fragment Dissociation.
- Frequency adaptive metadynamics for the calculation of rare-event kinetics.
- A Multiscale Simulation Approach to Modeling Drug-Protein Binding Kinetics.
- Computational Study of Protein-Ligand Unbinding for Enzyme Engineering.
- How and when does an anticancer drug leave its binding site?
- Unbinding Kinetics of a p38 MAP Kinase Type II Inhibitor from Metadynamics Simulations.
- Metadynamics Simulations Distinguish Short- and Long-Residence-Time Inhibitors of Cyclin-Dependent Kinase 8.
- Characterizing Drug-Target Residence Time with Metadynamics: How To Achieve Dissociation Rate Efficiently without Losing Accuracy against Time-Consuming Approaches.
- Kinetics of protein-ligand unbinding: Predicting pathways, rates, and rate-limiting steps.
- Decoding the Role of Water Dynamics in Ligand-Protein Unbinding: CRF1R as a Test Case.
Metadynamics was also used in the following examples:
- Transient States and Barriers from Molecular Simulations and the Milestoning Theory: Kinetics in Ligand-Protein Recognition and Compound Design.
- Enhanced Sampling Simulations of Ligand Unbinding Kinetics Controlled by Protein Conformational Changes.
- Substrate binding mechanism of HIV-1 protease from explicit-solvent atomistic simulations.