BKiT
Availablilty
URL: https://github.com/chang-group/BKiT
Licence: Free
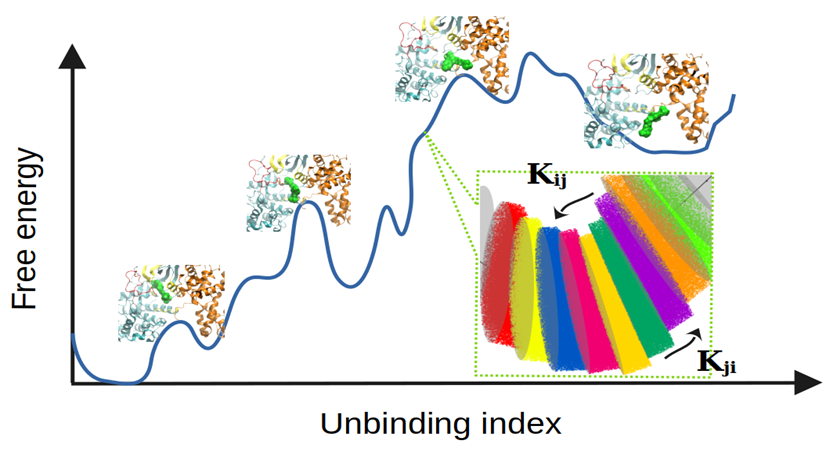
Binding Kinetics Toolkit (BKiT) is a computational toolkit which contains several utilities able to analyze molecular dynamic (MD) simulations to extract thermodynamic and kinetic information. Through the use of principal component space, ligand unbinding or protein conformational transitions can easily be partitioned into microstates or milestones allowing the users to visualize molecular motions for detailed analysis. This program can also generate input files to run many short classical MD simulations. With the use of milestoning theory, free energy profiles are constructed, and residence times are estimated. The source code is written in Python and allows users to tailor the toolkit to their specific needs.